Sparse Gaussian Process Regression¶
Let’s set some setting for this Jupyter Notebook.
In [2]:
%matplotlib inline
from warnings import filterwarnings
filterwarnings("ignore")
import os
os.environ['MKL_THREADING_LAYER'] = 'GNU'
os.environ['THEANO_FLAGS'] = 'device=cpu'
import numpy as np
import pandas as pd
import pymc3 as pm
import seaborn as sns
import matplotlib.pyplot as plt
np.random.seed(12345)
rc = {'xtick.labelsize': 20, 'ytick.labelsize': 20, 'axes.labelsize': 20, 'font.size': 20,
'legend.fontsize': 12.0, 'axes.titlesize': 10, "figure.figsize": [12, 6]}
sns.set(rc = rc)
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
Now, let’s import the SparseGaussianProcessRegression algorithm from
the pymc-learn package.
In [3]:
import pmlearn
from pmlearn.gaussian_process import SparseGaussianProcessRegressor
print('Running on pymc-learn v{}'.format(pmlearn.__version__))
Running on pymc-learn v0.0.1.rc0
Step 1: Prepare the data¶
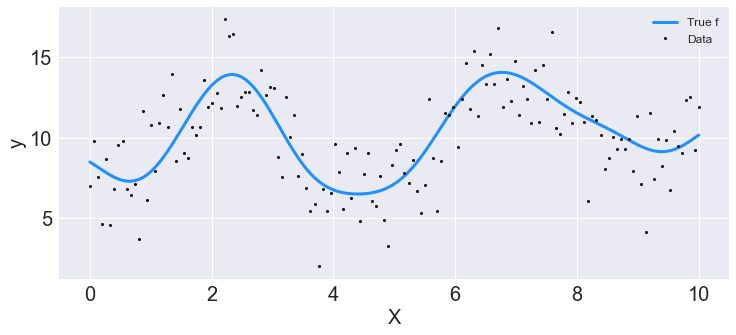
Generate synthetic data.
In [4]:
n = 150 # The number of data points
X = np.linspace(start = 0, stop = 10, num = n)[:, None] # The inputs to the GP, they must be arranged as a column vector
# Define the true covariance function and its parameters
length_scale_true = 1.0
signal_variance_true = 3.0
cov_func = signal_variance_true**2 * pm.gp.cov.ExpQuad(1, length_scale_true)
# A mean function that is zero everywhere
mean_func = pm.gp.mean.Constant(10)
# The latent function values are one sample from a multivariate normal
# Note that we have to call `eval()` because PyMC3 built on top of Theano
f_true = np.random.multivariate_normal(mean_func(X).eval(),
cov_func(X).eval() + 1e-8*np.eye(n), 1).flatten()
# The observed data is the latent function plus a small amount of Gaussian distributed noise
# The standard deviation of the noise is `sigma`
noise_variance_true = 2.0
y = f_true + noise_variance_true * np.random.randn(n)
## Plot the data and the unobserved latent function
fig = plt.figure(figsize=(12,5))
ax = fig.gca()
ax.plot(X, f_true, "dodgerblue", lw=3, label="True f");
ax.plot(X, y, 'ok', ms=3, label="Data");
ax.set_xlabel("X"); ax.set_ylabel("y"); plt.legend();
In [5]:
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
Step 2: Instantiate a model¶
In [6]:
model = SparseGaussianProcessRegressor()
Step 3: Perform Inference¶
In [7]:
model.fit(X_train, y_train)
Average Loss = -4,669.3: 100%|██████████| 200000/200000 [06:12<00:00, 536.41it/s]
Finished [100%]: Average Loss = -4,669.6
Out[7]:
SparseGaussianProcessRegressor(prior_mean=0.0)
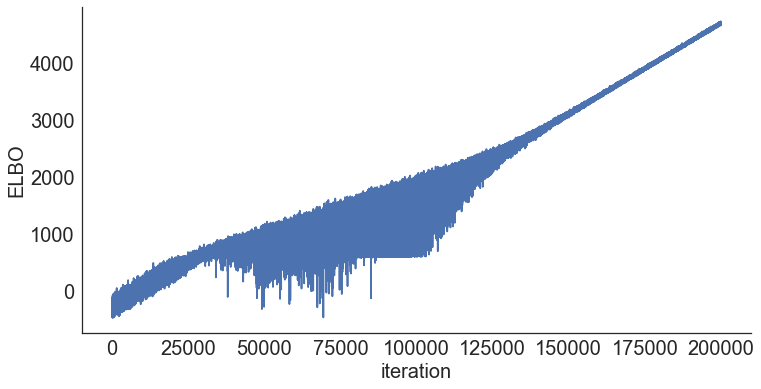
Step 4: Diagnose convergence¶
In [8]:
model.plot_elbo()
In [8]:
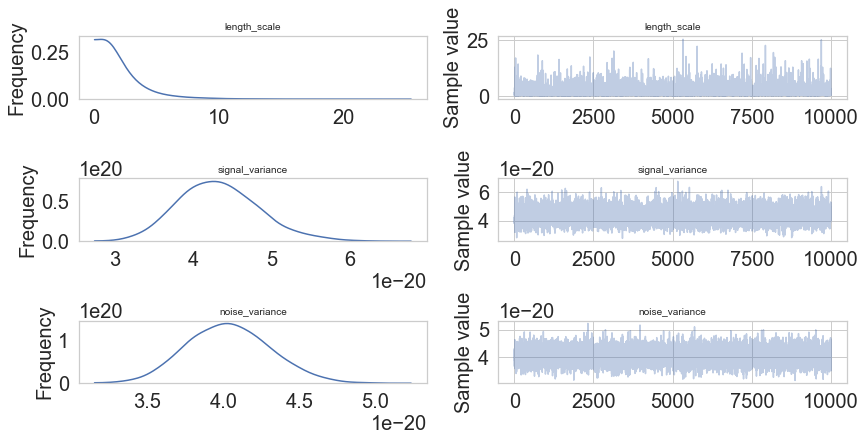
pm.traceplot(model.trace);
In [9]:
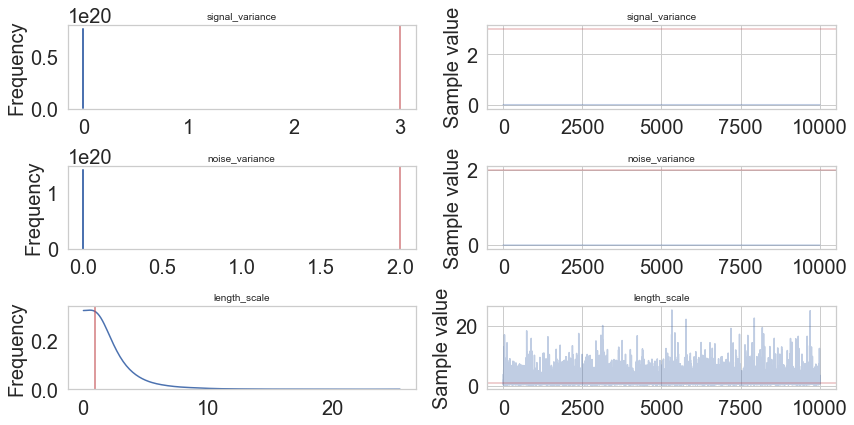
pm.traceplot(model.trace, lines = {"signal_variance": signal_variance_true,
"noise_variance": noise_variance_true,
"length_scale": length_scale_true},
varnames=["signal_variance", "noise_variance", "length_scale"]);
In [11]:
pm.forestplot(model.trace, varnames=["signal_variance", "noise_variance", "length_scale"]);
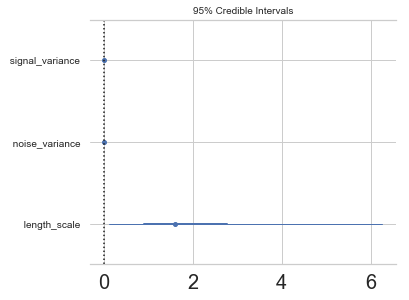
Step 5: Critize the model¶
In [12]:
pm.summary(model.trace, varnames=["signal_variance", "length_scale", "noise_variance"])
Out[12]:
| mean | sd | mc_error | hpd_2.5 | hpd_97.5 | |
|---|---|---|---|---|---|
| signal_variance__0 | 2.266128e-20 | 3.632205e-21 | 3.731201e-23 | 1.578536e-20 | 2.999736e-20 |
| length_scale__0_0 | 2.233311e+00 | 2.182089e+00 | 2.042519e-02 | 1.187429e-01 | 6.257235e+00 |
| noise_variance__0 | 2.216509e-20 | 3.491209e-21 | 3.041214e-23 | 1.571128e-20 | 2.914439e-20 |
In [13]:
pm.plot_posterior(model.trace, varnames=["signal_variance", "noise_variance", "length_scale"],
figsize = [14, 8]);
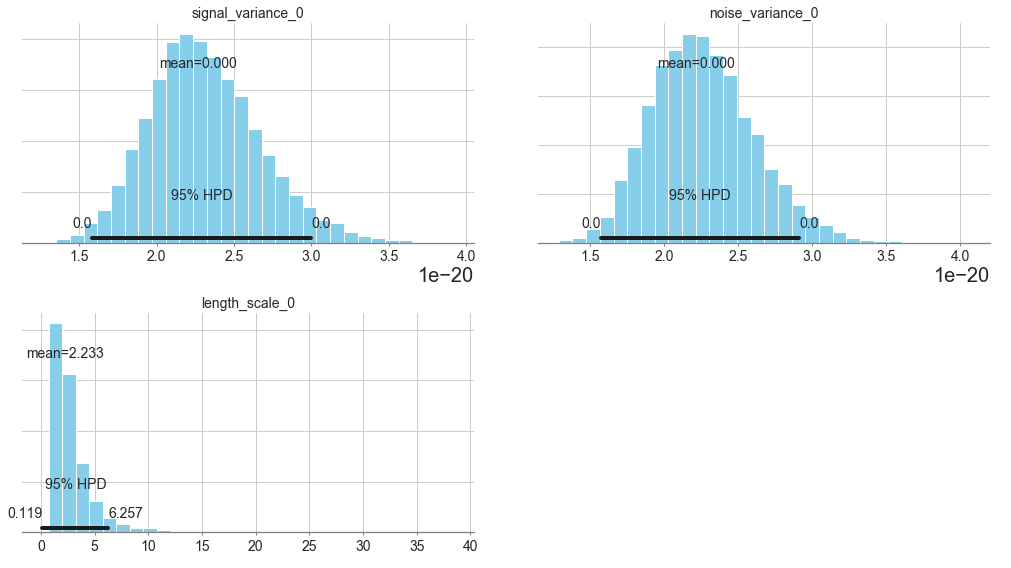
In [8]:
# collect the results into a pandas dataframe to display
# "mp" stands for marginal posterior
pd.DataFrame({"Parameter": ["length_scale", "signal_variance", "noise_variance"],
"Predicted Mean Value": [float(model.trace["length_scale"].mean(axis=0)),
float(model.trace["signal_variance"].mean(axis=0)),
float(model.trace["noise_variance"].mean(axis=0))],
"True value": [length_scale_true, signal_variance_true, noise_variance_true]})
Out[8]:
| Parameter | Predicted Mean Value | True value | |
|---|---|---|---|
| 0 | length_scale | 2.182521 | 1.0 |
| 1 | signal_variance | 9.261435 | 3.0 |
| 2 | noise_variance | 0.002241 | 2.0 |
Step 6: Use the model for prediction¶
In [ ]:
y_predict1 = model.predict(X_test)
In [ ]:
y_predict1
In [ ]:
model.score(X_test, y_test)
In [ ]:
model.save('pickle_jar/sgpr')
Use already trained model for prediction¶
In [ ]:
model_new = SparseGaussianProcessRegressor()
model_new.load('pickle_jar/sgpr')
model_new.score(X_test, y_test)
Multiple Features¶
In [ ]:
num_pred = 2
X = np.random.randn(1000, num_pred)
noise = 2 * np.random.randn(1000,)
Y = X.dot(np.array([4, 5])) + 3 + noise
In [ ]:
y = np.squeeze(Y)
In [ ]:
model_big = SparseGaussianProcessRegressor()
In [ ]:
model_big.fit(X, y, inference_args={"n" : 1000})
In [ ]:
pm.summary(model_big.trace, varnames=["signal_variance", "length_scale", "noise_variance"])
MCMC¶
In [8]:
model2 = SparseGaussianProcessRegressor()
model2.fit(X_train, y_train, inference_type='nuts')
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [f_rotated_, noise_variance_log__, signal_variance_log__, length_scale_log__]
100%|██████████| 1500/1500 [00:24<00:00, 60.20it/s]
There were 92 divergences after tuning. Increase `target_accept` or reparameterize.
There were 88 divergences after tuning. Increase `target_accept` or reparameterize.
There were 87 divergences after tuning. Increase `target_accept` or reparameterize.
There were 40 divergences after tuning. Increase `target_accept` or reparameterize.
The number of effective samples is smaller than 10% for some parameters.
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [f_rotated_, noise_variance_log__, signal_variance_log__, length_scale_log__]
100%|██████████| 4000/4000 [11:41<00:00, 5.70it/s]
There were 4 divergences after tuning. Increase `target_accept` or reparameterize.
There were 2 divergences after tuning. Increase `target_accept` or reparameterize.
There were 1 divergences after tuning. Increase `target_accept` or reparameterize.
There were 1 divergences after tuning. Increase `target_accept` or reparameterize.
Out[8]:
GaussianProcessRegression()
In [18]:
pm.traceplot(model2.trace, lines = {"signal_variance": signal_variance_true,
"noise_variance": noise_variance_true,
"length_scale": length_scale_true},
varnames=["signal_variance", "noise_variance", "length_scale"]);
In [19]:
pm.gelman_rubin(model2.trace, varnames=["signal_variance", "noise_variance", "length_scale"])
Out[19]:
{'signal_variance': array([ 1.00134827]),
'noise_variance': array([ 0.99982997]),
'length_scale': array([[ 0.9997668]])}
In [22]:
pm.energyplot(model2.trace);
In [21]:
pm.forestplot(model2.trace, varnames=["signal_variance", "noise_variance", "length_scale"]);
In [10]:
pm.summary(model2.trace, varnames=["signal_variance", "length_scale", "noise_variance"])
Out[10]:
| mean | sd | mc_error | hpd_2.5 | hpd_97.5 | n_eff | Rhat | |
|---|---|---|---|---|---|---|---|
| signal_variance__0 | 3.354521 | 5.059969 | 0.072611 | 0.004388 | 10.494186 | 3949.565916 | 1.001348 |
| length_scale__0_0 | 2.004001 | 1.446405 | 0.013286 | 0.033810 | 4.790922 | 12131.009636 | 0.999767 |
| noise_variance__0 | 2.544328 | 0.264045 | 0.003021 | 2.074630 | 3.086981 | 8803.802924 | 0.999830 |
In [11]:
# collect the results into a pandas dataframe to display
# "mp" stands for marginal posterior
pd.DataFrame({"Parameter": ["length_scale", "signal_variance", "noise_variance"],
"Predicted Mean Value": [float(model2.trace["length_scale"].mean(axis=0)),
float(model2.trace["signal_variance"].mean(axis=0)),
float(model2.trace["noise_variance"].mean(axis=0))],
"True value": [length_scale_true, signal_variance_true, noise_variance_true]})
Out[11]:
| Parameter | Predicted Mean Value | True value | |
|---|---|---|---|
| 0 | length_scale | 2.004001 | 1.0 |
| 1 | signal_variance | 3.354521 | 3.0 |
| 2 | noise_variance | 2.544328 | 2.0 |
In [12]:
pm.plot_posterior(model2.trace, varnames=["signal_variance", "noise_variance", "length_scale"],
figsize = [14, 8]);
In [28]:
y_predict2 = model2.predict(X_test)
100%|██████████| 2000/2000 [00:01<00:00, 1332.79it/s]
In [29]:
y_predict2
Out[29]:
array([ 0.71831924, 0.70266214, 0.74034292, 0.73223746, 0.76798942,
0.78039904, 0.80198739, 0.77559783, 0.74532885, 0.75839183,
0.69163726, 0.6490964 , 0.71534946, 0.65845406, 0.66052402,
0.80801464, 0.69148553, 0.61070685, 0.69928683, 0.75866764,
0.6620472 , 0.73977574, 0.70854909, 0.70340364, 0.70960481,
0.69097856, 0.69340258, 0.72408786, 0.81266196, 0.79486012,
0.72997809, 0.66805751, 0.72690218, 0.71025724, 0.72545681,
0.69062513, 0.75047548, 0.64446808, 0.78133024, 0.69365793,
0.78675961, 0.7909775 , 0.66224847, 0.67357815, 0.82613138,
0.76196312, 0.76742 , 0.67757641, 0.67067013, 0.70072039])
In [30]:
model2.score(X_test, y_test)
100%|██████████| 2000/2000 [00:01<00:00, 1464.52it/s]
Out[30]:
-0.011313490552906202
In [ ]:
model2.save('pickle_jar/')
model2_new = SparseGaussianProcessRegressor()
model2_new.load('pickle_jar/')
model2_new.score(X_test, y_test)