Student’s T Process Regression¶
Let’s set some setting for this Jupyter Notebook.
In [2]:
%matplotlib inline
from warnings import filterwarnings
filterwarnings("ignore")
import os
os.environ['MKL_THREADING_LAYER'] = 'GNU'
os.environ['THEANO_FLAGS'] = 'device=cpu'
import numpy as np
import pandas as pd
import pymc3 as pm
import seaborn as sns
import matplotlib.pyplot as plt
np.random.seed(12345)
rc = {'xtick.labelsize': 20, 'ytick.labelsize': 20, 'axes.labelsize': 20, 'font.size': 20,
'legend.fontsize': 12.0, 'axes.titlesize': 10, "figure.figsize": [12, 6]}
sns.set(rc = rc)
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
Now, let’s import the StudentsTProcessRegression algorithm from the
pymc-learn package.
In [3]:
import pmlearn
from pmlearn.gaussian_process import StudentsTProcessRegressor
print('Running on pymc-learn v{}'.format(pmlearn.__version__))
Running on pymc-learn v0.0.1.rc0
Step 1: Prepare the data¶
Generate synthetic data.
In [4]:
n = 150 # The number of data points
X = np.linspace(start = 0, stop = 10, num = n)[:, None] # The inputs to the GP, they must be arranged as a column vector
# Define the true covariance function and its parameters
length_scale_true = 1.0
signal_variance_true = 3.0
cov_func = signal_variance_true**2 * pm.gp.cov.ExpQuad(1, length_scale_true)
# A mean function that is zero everywhere
mean_func = pm.gp.mean.Zero()
# The latent function values are one sample from a multivariate normal
# Note that we have to call `eval()` because PyMC3 built on top of Theano
f_true = np.random.multivariate_normal(mean_func(X).eval(),
cov_func(X).eval() + 1e-8*np.eye(n), 1).flatten()
# The observed data is the latent function plus a small amount of T distributed noise
# The standard deviation of the noise is `sigma`, and the degrees of freedom is `nu`
noise_variance_true = 2.0
degrees_of_freedom_true = 3.0
y = f_true + noise_variance_true * np.random.standard_t(degrees_of_freedom_true, size=n)
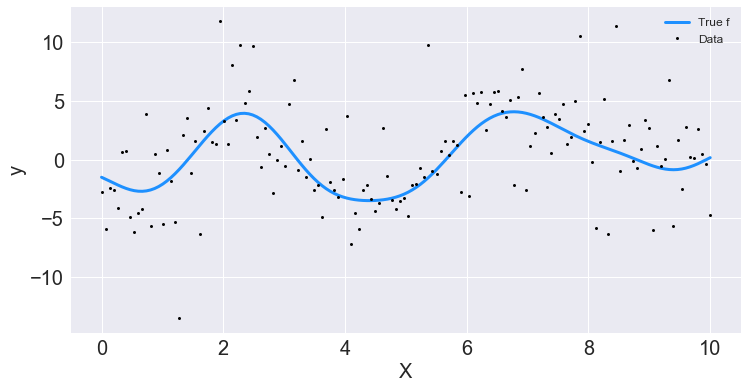
## Plot the data and the unobserved latent function
fig, ax = plt.subplots()
ax.plot(X, f_true, "dodgerblue", lw=3, label="True f");
ax.plot(X, y, 'ok', ms=3, label="Data");
ax.set_xlabel("X"); ax.set_ylabel("y"); plt.legend();
In [5]:
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
Step 2: Instantiate a model¶
In [6]:
model = StudentsTProcessRegressor()
Step 3: Perform Inference¶
In [7]:
model.fit(X_train, y_train)
Average Loss = 303.15: 100%|██████████| 200000/200000 [06:37<00:00, 503.33it/s]
Finished [100%]: Average Loss = 303.15
Out[7]:
StudentsTProcessRegressor(prior_mean=0.0)
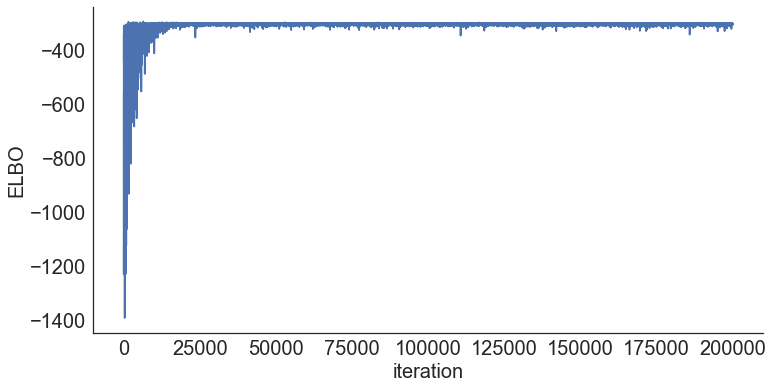
Step 4: Diagnose convergence¶
In [8]:
model.plot_elbo()
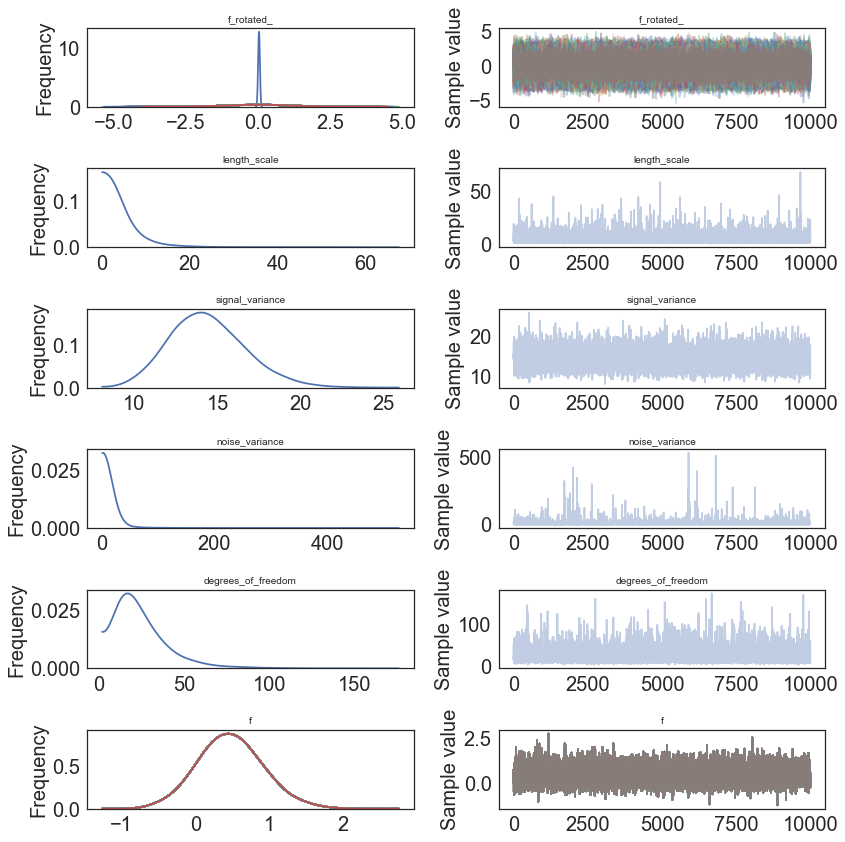
In [9]:
pm.traceplot(model.trace);
In [10]:
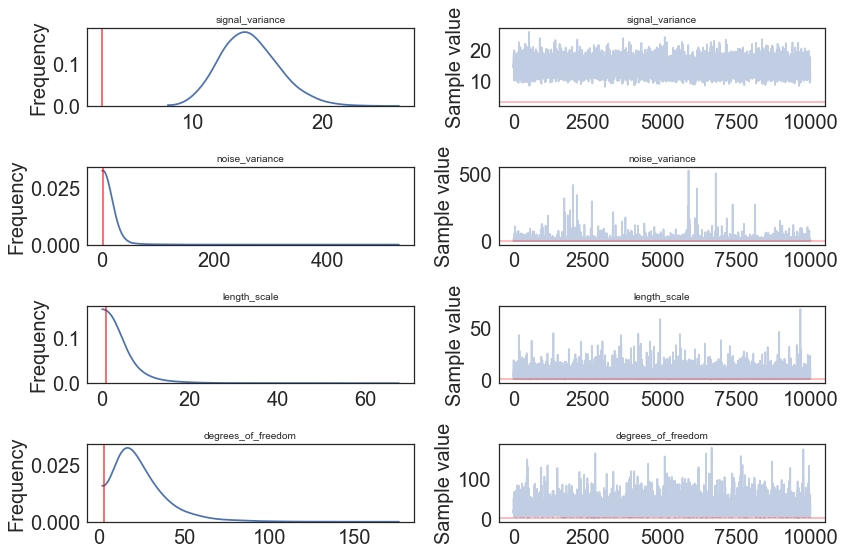
pm.traceplot(model.trace, lines = {"signal_variance": signal_variance_true,
"noise_variance": noise_variance_true,
"length_scale": length_scale_true,
"degrees_of_freedom": degrees_of_freedom_true},
varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"]);
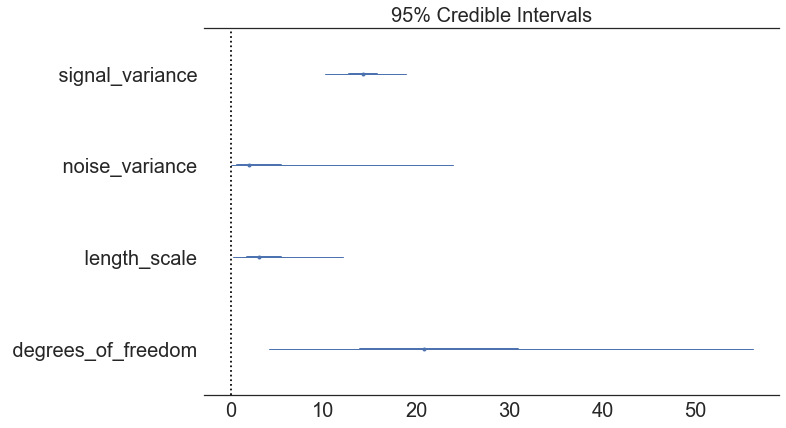
In [11]:
pm.forestplot(model.trace, varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"]);
Step 5: Criticize the model¶
In [12]:
pm.summary(model.trace, varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"])
Out[12]:
| mean | sd | mc_error | hpd_2.5 | hpd_97.5 | |
|---|---|---|---|---|---|
| signal_variance__0 | 14.369054 | 2.254097 | 0.021957 | 10.170910 | 18.868740 |
| noise_variance__0 | 6.182008 | 17.330061 | 0.187540 | 0.006105 | 23.940172 |
| length_scale__0_0 | 4.348170 | 4.162548 | 0.042757 | 0.319785 | 12.072470 |
| degrees_of_freedom__0 | 24.762167 | 16.178113 | 0.149632 | 4.172859 | 56.188869 |
In [13]:
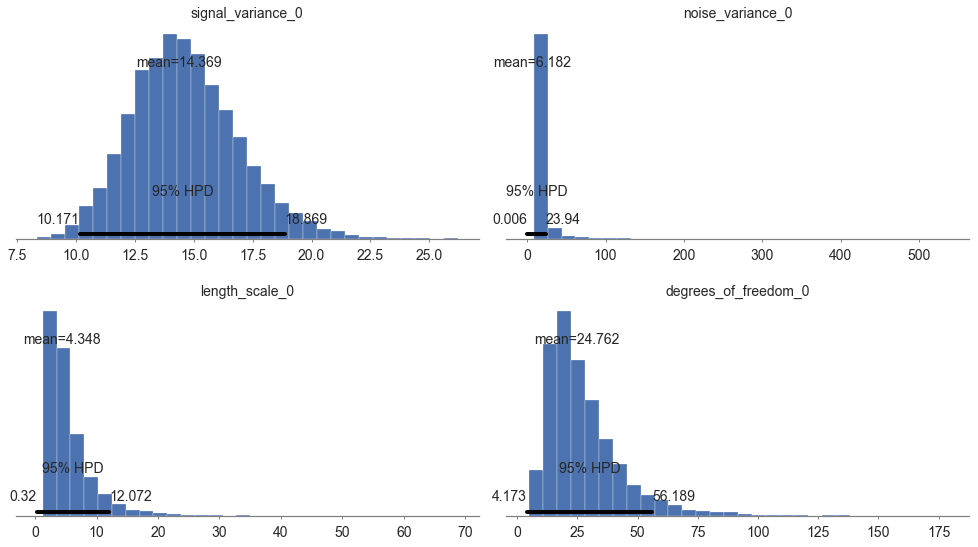
pm.plot_posterior(model.trace, varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"],
figsize = [14, 8]);
In [14]:
# collect the results into a pandas dataframe to display
# "mp" stands for marginal posterior
pd.DataFrame({"Parameter": ["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"],
"Predicted Mean Value": [float(model.trace["length_scale"].mean(axis=0)),
float(model.trace["signal_variance"].mean(axis=0)),
float(model.trace["noise_variance"].mean(axis=0)),
float(model.trace["degrees_of_freedom"].mean(axis=0))],
"True value": [length_scale_true, signal_variance_true,
noise_variance_true, degrees_of_freedom_true]})
Out[14]:
| Parameter | Predicted Mean Value | True value | |
|---|---|---|---|
| 0 | signal_variance | 4.348170 | 1.0 |
| 1 | noise_variance | 14.369054 | 3.0 |
| 2 | length_scale | 6.182008 | 2.0 |
| 3 | degrees_of_freedom | 24.762167 | 3.0 |
Step 6: Use the model for prediction¶
In [15]:
y_predict1 = model.predict(X_test)
100%|██████████| 2000/2000 [00:08<00:00, 247.33it/s]
In [16]:
y_predict1
Out[16]:
array([0.52060618, 0.20859641, 0.32341845, 0.71517795, 0.12535947,
0.10130519, 0.13356278, 0.48476055, 0.33239652, 0.05354277,
0.3221012 , 0.27747592, 0.33224296, 0.16754793, 0.70514462,
0.37293254, 0.38020924, 0.65038549, 0.34252208, 0.38382534,
0.15502318, 0.37618247, 0.58213956, 0.63244638, 0.27682323,
0.17309081, 0.11088147, 0.38385589, 0.05206571, 0.33370627,
0.0590494 , 0.21805391, 0.24068462, 0.14248978, 0.16113507,
0.6395228 , 0.13902426, 0.29770677, 0.24498306, 0.18377858,
0.12288624, 0.35066241, 0.25833606, 0.70100999, 0.66802676])
In [24]:
model.score(X_test, y_test)
In [26]:
model.save('pickle_jar/spr')
Use already trained model for prediction¶
In [27]:
model_new = StudentsTProcessRegressor()
model_new.load('pickle_jar/spr')
model_new.score(X_test, y_test)
100%|██████████| 2000/2000 [00:01<00:00, 1201.17it/s]
Out[27]:
-0.09713232621579238
Multiple Features¶
In [34]:
num_pred = 2
X = np.random.randn(1000, num_pred)
noise = 2 * np.random.randn(1000,)
Y = X.dot(np.array([4, 5])) + 3 + noise
In [35]:
y = np.squeeze(Y)
In [36]:
model_big = StudentsTProcessRegressor()
In [37]:
model_big.fit(X, y, inference_args={"n" : 1000})
Average Loss = 6,129.9: 100%|██████████| 1000/1000 [03:13<00:00, 5.16it/s]
Finished [100%]: Average Loss = 6,118.9
Out[37]:
StudentsTProcessRegression()
In [38]:
pm.summary(model_big.trace, varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"])
Out[38]:
| mean | sd | mc_error | hpd_2.5 | hpd_97.5 | |
|---|---|---|---|---|---|
| signal_variance__0 | 7.029373 | 4.739398 | 0.049285 | 0.968977 | 16.254415 |
| noise_variance__0 | 7.163999 | 7.395869 | 0.075956 | 0.337118 | 20.242566 |
| length_scale__0_0 | 2.451322 | 1.983714 | 0.022889 | 0.256500 | 6.175940 |
| length_scale__0_1 | 2.466894 | 2.009942 | 0.021930 | 0.196610 | 6.184087 |
| degrees_of_freedom__0 | 19.622088 | 15.718934 | 0.147647 | 2.390572 | 49.401395 |
MCMC¶
In [8]:
model2 = StudentsTProcessRegressor()
model2.fit(X_train, y_train, inference_type='nuts')
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [f_rotated_, degrees_of_freedom_log__, noise_variance_log__, signal_variance_log__, length_scale_log__]
100%|██████████| 2500/2500 [03:33<00:00, 11.70it/s]
Out[8]:
StudentsTProcessRegression()
In [13]:
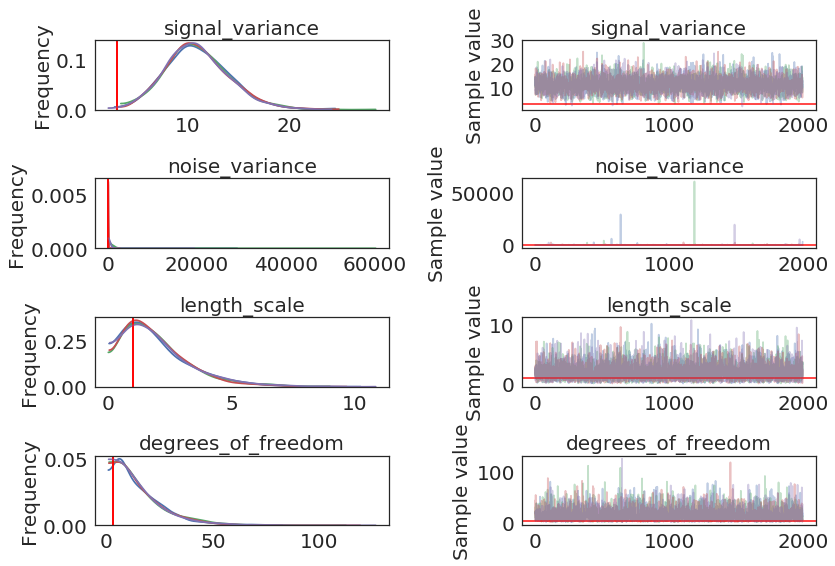
pm.traceplot(model2.trace, lines = {"signal_variance": signal_variance_true,
"noise_variance": noise_variance_true,
"length_scale": length_scale_true,
"degrees_of_freedom": degrees_of_freedom_true},
varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"]);
In [14]:
pm.gelman_rubin(model2.trace, varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"])
Out[14]:
{'degrees_of_freedom': array([1.00019487]),
'length_scale': array([[1.00008203]]),
'noise_variance': array([0.99986753]),
'signal_variance': array([0.99999439])}
In [15]:
pm.energyplot(SPR2.trace);
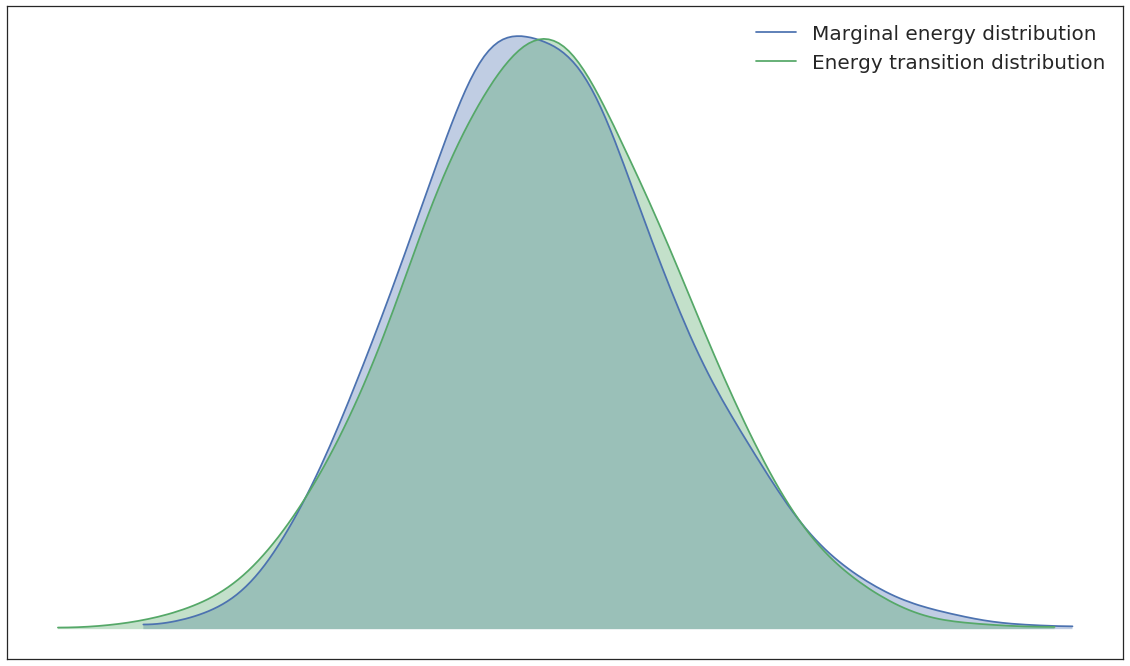
In [16]:
pm.forestplot(model2.trace, varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"]);
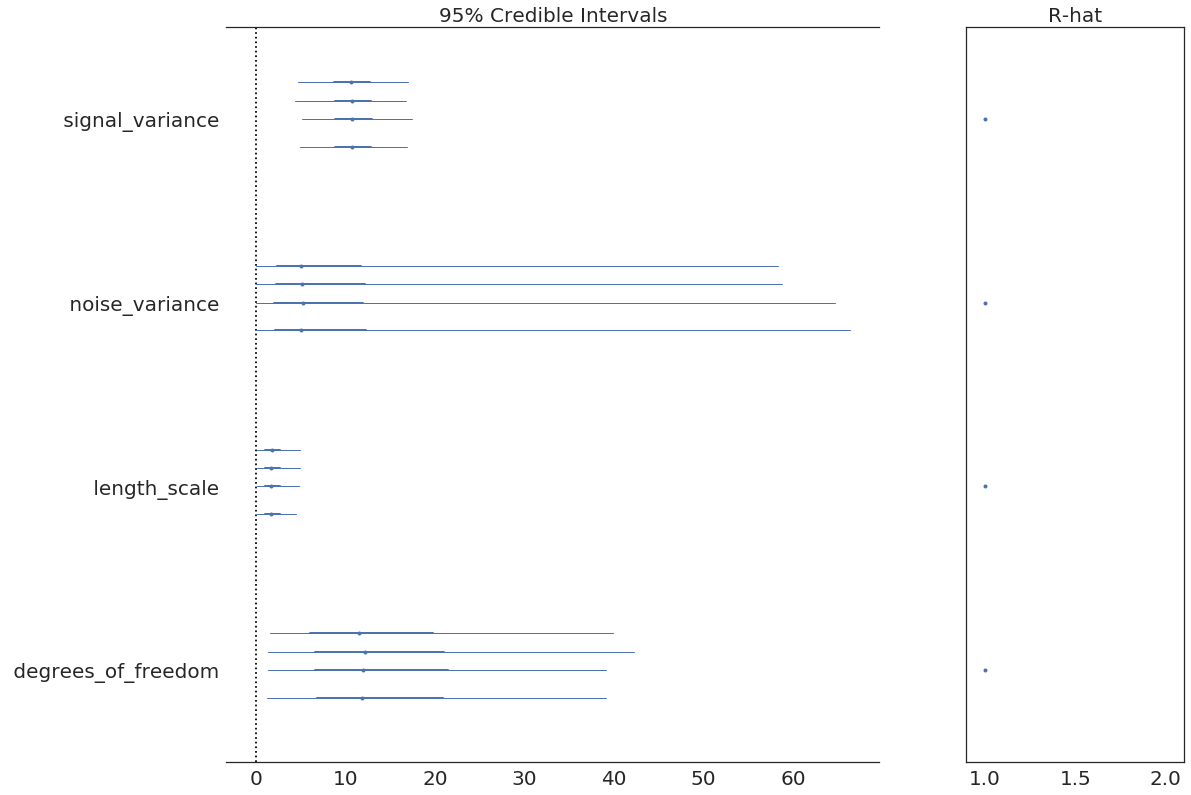
In [20]:
pm.summary(model2.trace, varnames=["signal_variance", "length_scale", "noise_variance"])
Out[20]:
| mean | sd | mc_error | hpd_2.5 | hpd_97.5 | n_eff | Rhat | |
|---|---|---|---|---|---|---|---|
| signal_variance__0 | 10.929238 | 3.144499 | 0.043256 | 4.628908 | 16.918602 | 4343.913263 | 0.999994 |
| length_scale__0_0 | 1.997809 | 1.397544 | 0.012827 | 0.061045 | 4.763012 | 11392.919787 | 1.000082 |
| noise_variance__0 | 36.850186 | 793.367946 | 9.474022 | 0.000534 | 63.332892 | 7491.212453 | 0.999868 |
In [21]:
# collect the results into a pandas dataframe to display
# "mp" stands for marginal posterior
pd.DataFrame({"Parameter": ["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"],
"Predicted Mean Value": [float(model2.trace["length_scale"].mean(axis=0)),
float(model2.trace["signal_variance"].mean(axis=0)),
float(model2.trace["noise_variance"].mean(axis=0)),
float(model2.trace["degrees_of_freedom"].mean(axis=0))],
"True value": [length_scale_true, signal_variance_true,
noise_variance_true, degrees_of_freedom_true]})
Out[21]:
| Parameter | Predicted Mean Value | True value | |
|---|---|---|---|
| 0 | signal_variance | 1.997809 | 1.0 |
| 1 | noise_variance | 10.929238 | 3.0 |
| 2 | length_scale | 36.850186 | 2.0 |
| 3 | degrees_of_freedom | 15.509439 | 3.0 |
In [22]:
pm.plot_posterior(model2.trace, varnames=["signal_variance", "noise_variance", "length_scale", "degrees_of_freedom"],
figsize = [14, 8]);
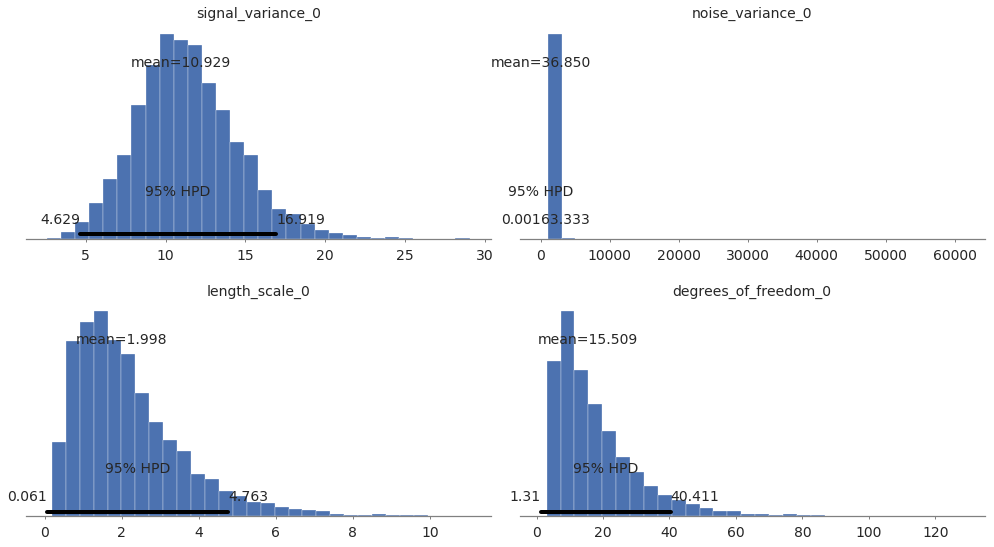
In [28]:
y_predict2 = model2.predict(X_test)
100%|██████████| 2000/2000 [00:01<00:00, 1174.91it/s]
In [29]:
y_predict2
Out[29]:
array([1.67834026, 1.64158368, 1.53728732, 1.56489496, 1.48686425,
1.48626043, 1.57801849, 1.5609818 , 1.57435388, 1.76800657,
1.56198154, 1.49355969, 1.72304612, 1.53818178, 1.68932836,
1.6059991 , 1.62152421, 1.50726857, 1.92453348, 1.61906672,
1.4703559 , 1.49874483, 1.63398678, 1.72795675, 1.62348916,
1.65877512, 1.78012082, 1.65401634, 1.47100635, 1.51878226,
1.53634253, 1.66642193, 1.5899548 , 1.62872435, 1.66256587,
1.67191658, 1.45945213, 1.43421284, 1.52586924, 1.56299994,
1.79883016, 1.6769178 , 1.52190602, 1.58302155, 1.44959024,
1.66465733, 1.5804623 , 1.62288222, 1.53714604, 1.80406125])
In [30]:
model2.score(X_test, y_test)
100%|██████████| 2000/2000 [00:01<00:00, 1254.66it/s]
Out[30]:
-0.0069721816446493
In [31]:
model2.save('pickle_jar/spr2')
model2_new = StudentsTProcessRegressor()
model2_new.load('pickle_jar/spr2')
model2_new.score(X_test, y_test)
100%|██████████| 2000/2000 [00:01<00:00, 1104.45it/s]
Out[31]:
0.0038373353227000306
Compare models¶
In [32]:
# plot the results
fig, ax = plt.subplots()
# plot the samples of the gp posterior
plt.plot(X_test, y_predict1, "r", lw=3, label="Predicted Mean Function")
plt.plot(X_train, f_true[100:], "dodgerblue", lw=3, label="True f");
plt.plot(X_test, y_test, 'ok', ms=3, alpha=0.5, label="Observed Test Data");
plt.xlabel("X")
plt.ylabel("True f(x)");
plt.title("ADVI: Conditional distribution of f_*, given f");
plt.legend();
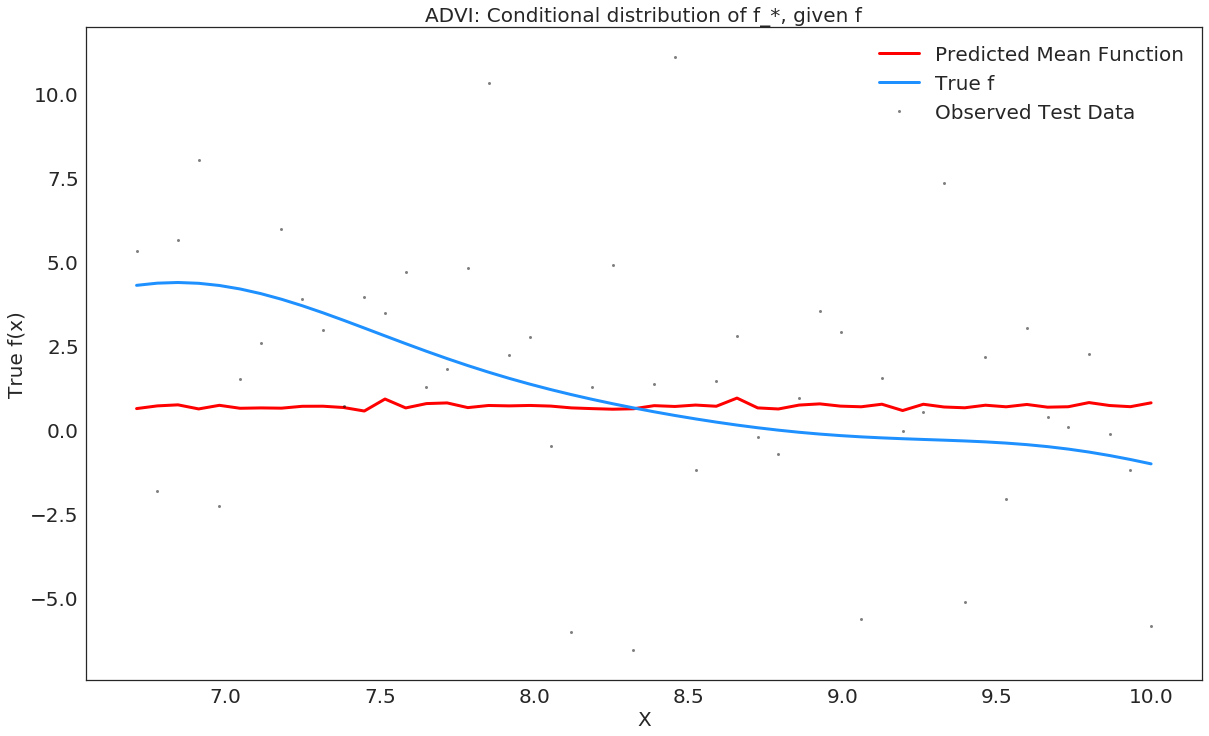
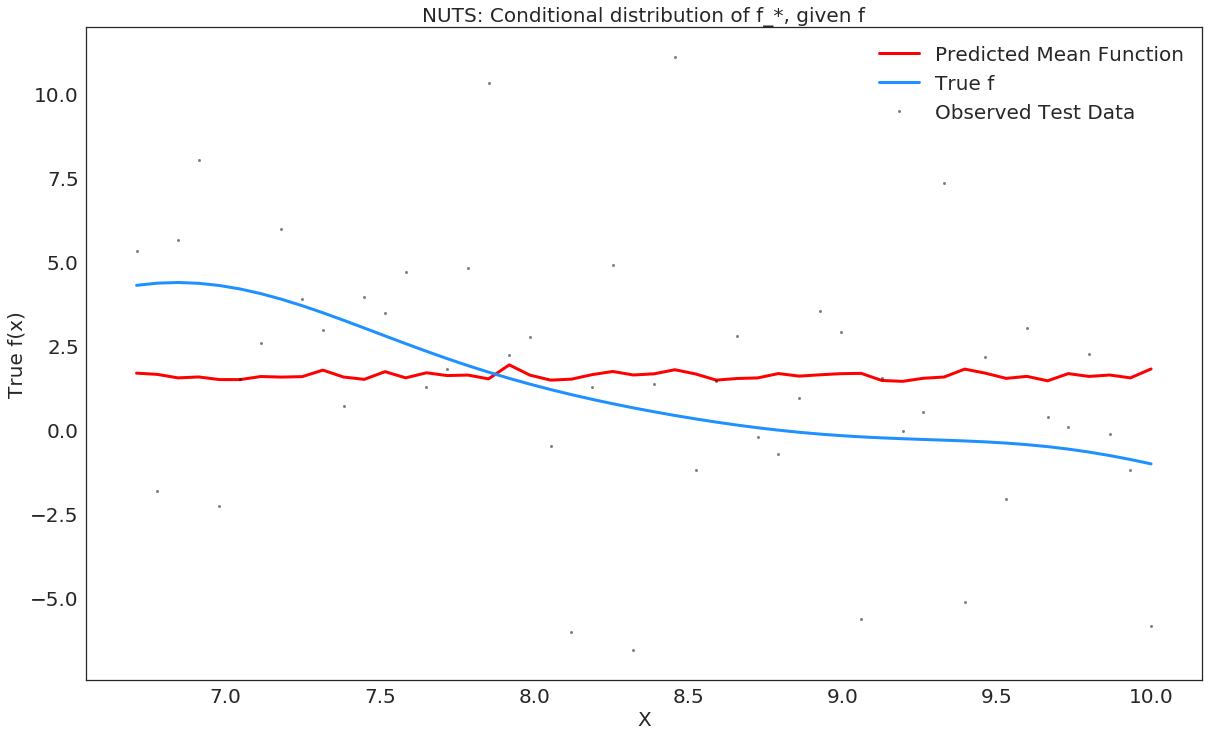
In [33]:
# plot the results
fig, ax = plt.subplots()
# plot the samples of the gp posterior
plt.plot(X_test, y_predict2, "r", lw=3, label="Predicted Mean Function")
plt.plot(X_train, f_true[100:], "dodgerblue", lw=3, label="True f");
plt.plot(X_test, y_test, 'ok', ms=3, alpha=0.5, label="Observed Test Data");
plt.xlabel("X")
plt.ylabel("True f(x)");
plt.title("NUTS: Conditional distribution of f_*, given f");
plt.legend();